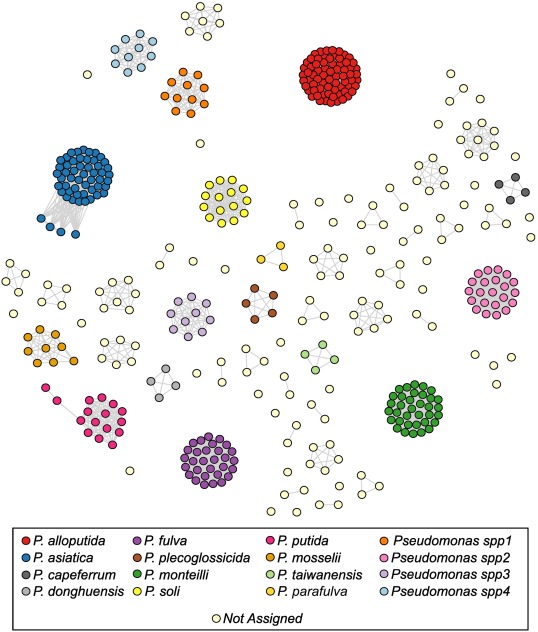
The Pseudomonas putida group comprises strains with biotechnological and clinical relevance. P. alloputida was proposed as a new species and highlighted the misclassification of P. putida. Nevertheless, the population structure of P. alloputida remained unexplored. We retrieved 11,025 Pseudomonas genomes and used P. alloputida Kh7T to delineate the species. The P. alloputida population structure comprises at least 7 clonal complexes (CCs). Clinical isolates are mainly found in CC4 and acquired resistance genes are present at low frequency in plasmids. Virulence profiles support the potential of CC7 members to outcompete other plant or human pathogens through a type VI secretion system. Finally, we found that horizontal gene transfer had an important role in shaping the ability of P. alloputida to bioremediate aromatic compounds such as toluene. Our results provide the grounds to understand P. alloputida genetic diversity and its potential for biotechnological applications.
Link of publication HERE
Publications 15
Research 2
 Posted on 08 Dec 2021
Posted on 08 Dec 2021
 Posted on 03 Nov 2021
Posted on 03 Nov 2021
New paper published! - Phylogenetic analysis and population structure of Pseudomonas alloputida
Posted on 03 Nov 2021 Posted on 03 Nov 2021
Posted on 03 Nov 2021