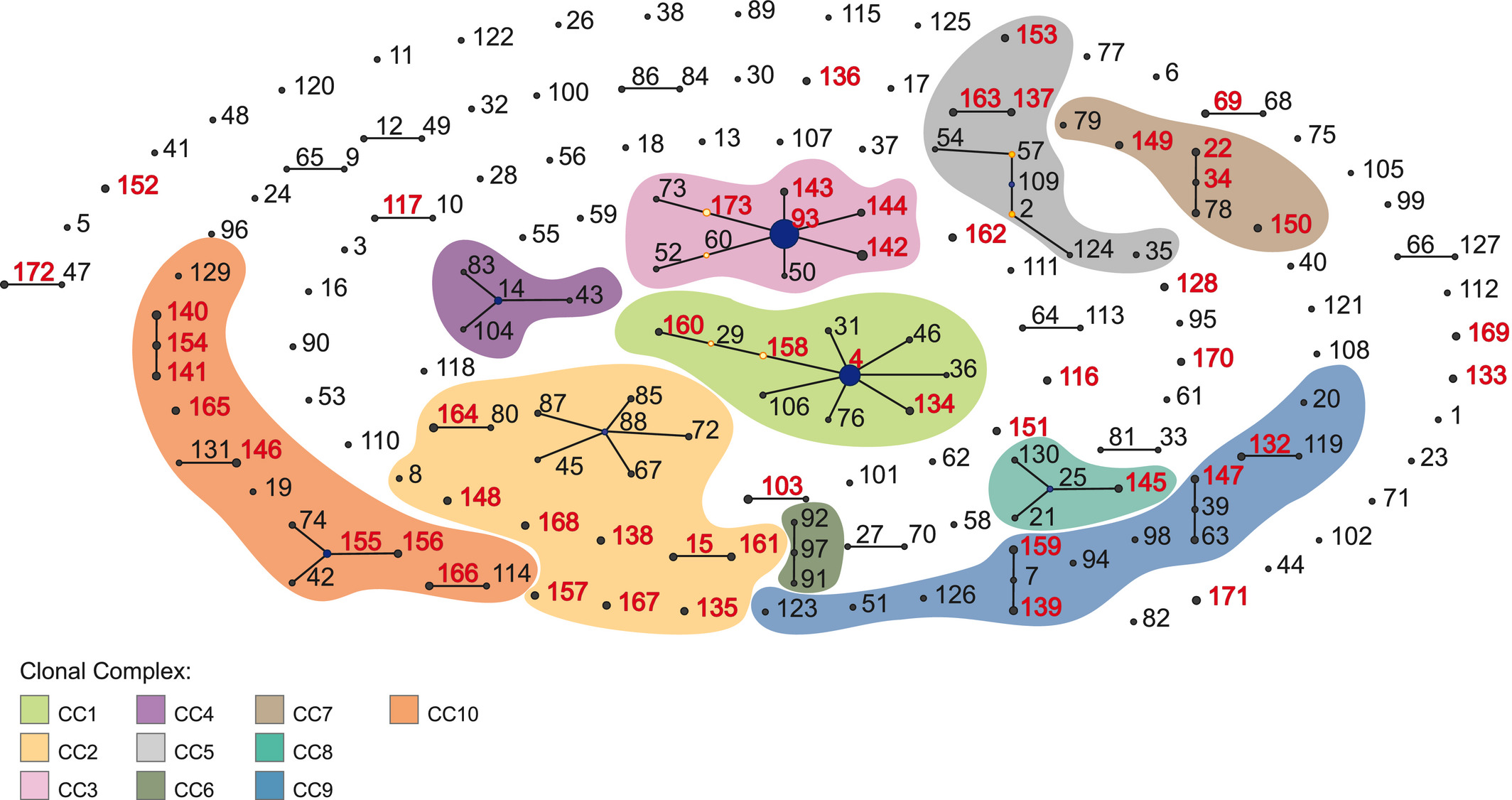
Klebsiella aerogenes is an important pathogen in healthcare‐associated infections. Nevertheless, in comparison to other clinically important pathogens, K. aerogenes population structure, genetic diversity, and pathogenicity remain poorly understood. Here, we elucidate K. aerogenes clonal complexes (CCs) and genomic features associated with resistance and virulence. We present a detailed description of the population structure of K. aerogenes based on 97 publicly available genomes by using both multilocus sequence typing and single‐nucleotide polymorphisms extracted from the core genome. We also assessed virulence and resistance profiles using Virulence Finder Database and Comprehensive Antibiotic Resistance Database, respectively. We show that K. aerogenes has an open pangenome and a large effective population size, which account for its high genomic diversity and support that negative selection prevents fixation of most deleterious alleles. The population is structured in at least 10 CCs, including two novel ones identified here, CC9 and CC10. The repertoires of resistance genes comprise a high number of antibiotic efflux proteins as well as narrow‐ and extended‐spectrum β‐lactamases. Regarding the population structure, we identified two clusters based on virulence profiles because of the presence of the toxin‐encoding clb operon and the siderophore production genes, irp and ybt. Notably, CC3 comprises the majority of K. aerogenes isolates associated with hospital outbreaks, emphasizing the importance of constant monitoring of this pathogen. Collectively, our results may provide a foundation for the development of new therapeutic and surveillance strategies worldwide.
Link of publication HERE
Publications 15
Research 2
 Posted on 08 Dec 2021
Posted on 08 Dec 2021
 Posted on 03 Nov 2021
Posted on 03 Nov 2021
New paper published! - Phylogenetic analysis and population structure of Pseudomonas alloputida
Posted on 03 Nov 2021 Posted on 03 Nov 2021
Posted on 03 Nov 2021